New DNAscent executable that measures inter-origin distance
- May 6
- 3 min read
We just realised DNAscent v4.2.1 which features the new executable meIODy (pronounced "melody") that measures inter-origin distance (IOD) from DNAscent data. This was developed by our undergraduate Part II Pathology student Olivia Mei, hence "meIODy" is a portmanteau of Mei, IOD, and melody.
IOD, the genomic distance between fired replication origins, is an important marker of DNA replication stress. When cells experience obstacles to fork progression (which may be from oncogene activation, nucleotide depletion, or exogenous DNA damage) they often compensate by firing additional replication origins. This reduces the spacing between fired origins, leading to a decrease in IOD.

IOD is traditionally measured with DNA fibre analysis. DNA fibres tend to have a length in excess of 300 kilobases (kb) [1] and most estimates of IOD in human cell lines lie around 100-150 kb [2,3] making the fibres long enough to capture IOD with relatively little bias caused by the distribution of fibre lengths. Measuring genome replication with Oxford Nanopore Technologies long-read DNA sequencing has always had a bit of a weakness in this area. The Oxford Nanopore reads, while very long, still only get to about an N50 of about 80 kb our hands [4]. This is considerably shorter than DNA fibres, and indeed shorter than the typical human cell IOD, making the reads too short to measure IOD without introducing a heavy bias that underestimates the real value. IOD was really the last area where fibre analysis had a big advantage over Oxford Nanopore-based methods, hence we were keen to innovate in this space to close the gap.

For us, the fun part about this project was that it didn't seem like it should be possible. How do you measure the distance between two things that your reads aren't long enough to capture? The answer turned out to be using clues left by how fork tracks were positioned on sequenced reads. meIODy extracts features of how much "empty read" lies ahead of, and behind, each fork track and then spins up a bunch of mathematical models that simulate what these features would look like at different IODs. Matching these features to the best model allows us to estimate a median IOD for each DNAscent sequencing run.

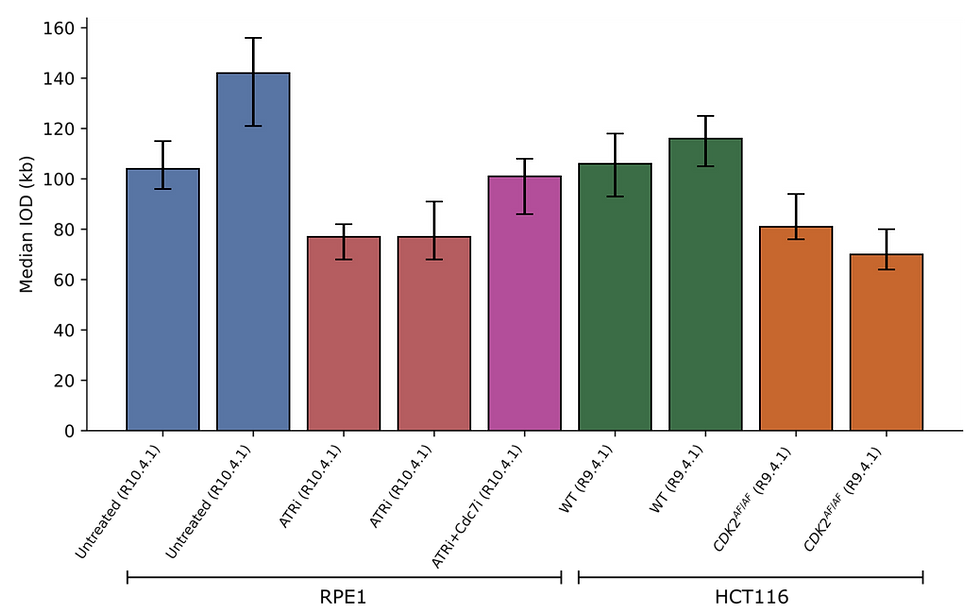
We found that IOD predictions from meIODy were consistent across biological replicates despite the sequencing runs having very different N50s. As expected, IOD decreased on treatment with ATRi in RPE1 cells [5]. In addition, the IOD decrease from ATRi treatment was rescued by inhibiting Cdc7. Estimates from meIODy on the IOD of HCT116 cells agreed with studies from DNA fibre [6] and introducing a mutation that prevents CDK2 inhibitory phosphorylation also showed a reduction in IOD.
One of our overarching design philosophies with DNAscent is that users should be able to squeeze more and more information out of their existing datasets as our algorithms continue to improve. In keeping with that ethos, meIODy works on DNAscent data from current R10.4.1 flow cells, but it is also back compatible with R9.4.1 DNAscent data, just like seeBreaks.
Olivia did a brilliant job of leading the algorithm development of meIODy, and it's absolutely remarkable that she drove this from idea-on-paper to software release in the span of a two-term undergraduate research project. We're also very grateful to our longstanding collaborators Mathew Jones, David Cullen, and Subash Rai from the University of Queensland with whom we are lucky to have worked on lots of interesting projects over the years. As ever, we like to get our new stuff into your hands as soon as possible, so we look forward to seeing what you do with it!
Team










